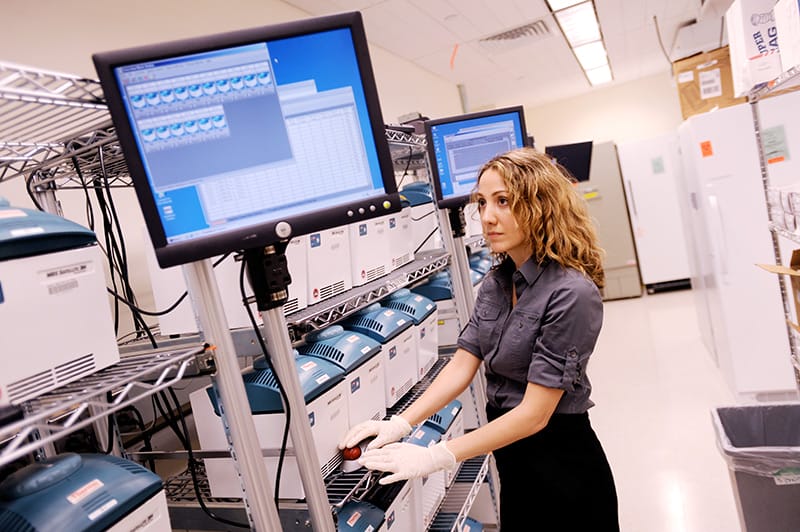

Long before the first case of Ebola was confirmed in the current outbreak, biologist Pardis Sabeti and her colleagues suspected that infectious diseases were already circulating, undetected. Pardis, an associate professor at Harvard University and winner of the 2014 Vilcek Prize for Creative Promise, had been working closely with the Kenema Government Hospital (KGH) in Sierra Leone to track and study another deadly hemorrhagic disease, Lassa fever, and Pardis was acutely aware of how difficult it would be to establish biologically secure environments to treat and control outbreaks of infectious diseases.
When the first case of Ebola was reported in neighboring Guinea, Pardis and her colleagues began routine testing of suspected patients. When the first case was confirmed in Sierra Leone, the team was approached by the country’s Ministry of Health and Sanitation to conduct genomic studies as a part of an emergency response to the outbreak.
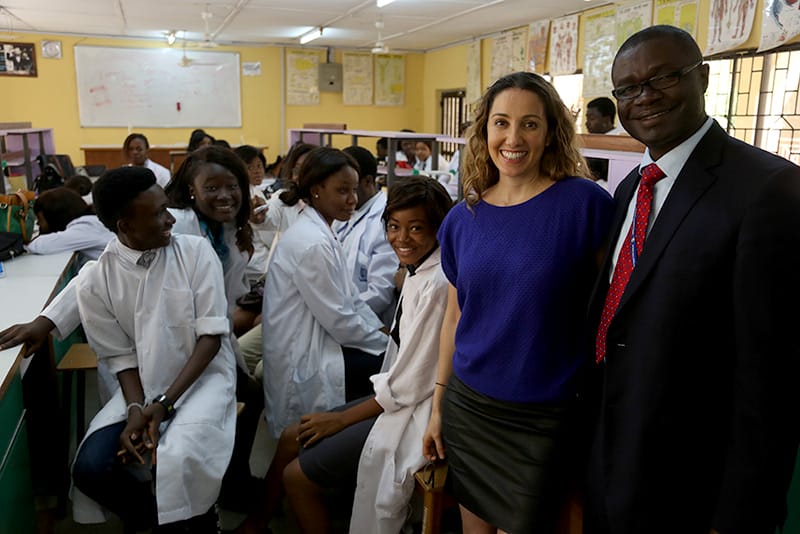

Pardis is exactly the person you’d want for the job. Combining Pardis’s experience in computational biology, evolutionary genetics, and infectious diseases, her lab has been receiving inactivated (read: non-infectious) samples of the virus to sequence. Sequences of the virus’s nucleic acid expose mutations in its genetic code, which are common during transmission.
Identification of the rapid accumulation of mutations gives greater insight into the outbreak. “It provides the evidence necessary to trace which viruses descended from other viruses and thus, which patients infected other patients. This allows us to take a much deeper look into the transmission of the virus—into the country, between towns, and between patients,” Pardis says. “Our findings were consistent with traditionally gathered epidemiology, but they also added much more detail than we would have had otherwise.” The mutations also illuminate transmission chains between disparate geographical locations.
In addition to tracking the outbreak, Pardis’s sequencing data is vital for developing diagnostic tools. “Ebola diagnosis is performed using a technique called polymerase chain reaction (PCR). PCR requires designing short sequences called primers, which can amplify a fragment of the Ebola virus’s genome if there is Ebola in the sample,” says Pardis. “The primer sequence should exactly match the viral sequence, so knowing the exact sequence of the viral strain that is currently infecting people is quite important for accurate diagnostics.”
Pardis’s lab has made the sequence data publicly available, which has generated a tremendous response from the scientific community. Researchers at the Scripps Research Institute have matched these sequences to antibodies in the ZMapp cocktail, an experimental therapeutic agent for Ebola, to see if the treatment would be effective against the West African variant of the virus. Other researchers are using the data to examine rare variants in the virus’s nucleic acid, which may not always be detected by traditional sequencing methods. “We can actually record a mutation rising to prevalence within a single host, or within a population, and this allows us to track how specific mutations arise, or what mutations might arise in the future,” says Pardis. “This is highly important for investigating vaccine and therapeutic targets.”
While scientific communities around the world have mobilized in response to the outbreak, much, much more could—and should—have been done to contain it. Pardis and her colleagues at KGH were aware of the dangers of a rampant infectious agent affecting the area long before the first reported case of Ebola. As the virus swept throughout the region and overwhelmed medical facilities, Sierra Leone lost many of its leading scientists and health care workers, many of whom had been Pardis’s colleagues.
“It did not have to be that way,” Pardis wrote in an opinion column for the New York Times dated Sept. 5, 2014. A stronger, coordinated effort to provide aid, support medical facilities, and train health professionals could have contained the outbreak. Educational and outreach services for local communities and secured delivery of food supplies could have lessened the impact on villages directly affected by the outbreak. “That missed opportunity has cost the lives of many people, including my good friends,” Pardis wrote.

The outbreak continues today, and Pardis is heartened by the fact that governments and international organizations are finally bringing the large surge of help that will be needed to stop the outbreaks raging in Sierra Leone, Liberia, and Guinea.
Pardis’s lab continues to work around the clock developing new diagnostic and sequencing tools, sending supplies and support to West African partners in Sierra Leone, Nigeria, and Senegal, and becoming involved in a number of initiatives to help the overall campaign.
Pardis and her lab have lost many beloved colleagues in this outbreak, so the fight for them is personal. In a rare break from their work, she and researchers from Nigeria and Senegal recently wrote and performed the song “One Truth” for those fallen, in which she states, “I’m here in this fight always.”
“We must end this outbreak, both to honor the memory of those who have died and to aid those still in the path of the virus. And then we must make sure that it never happens again.”